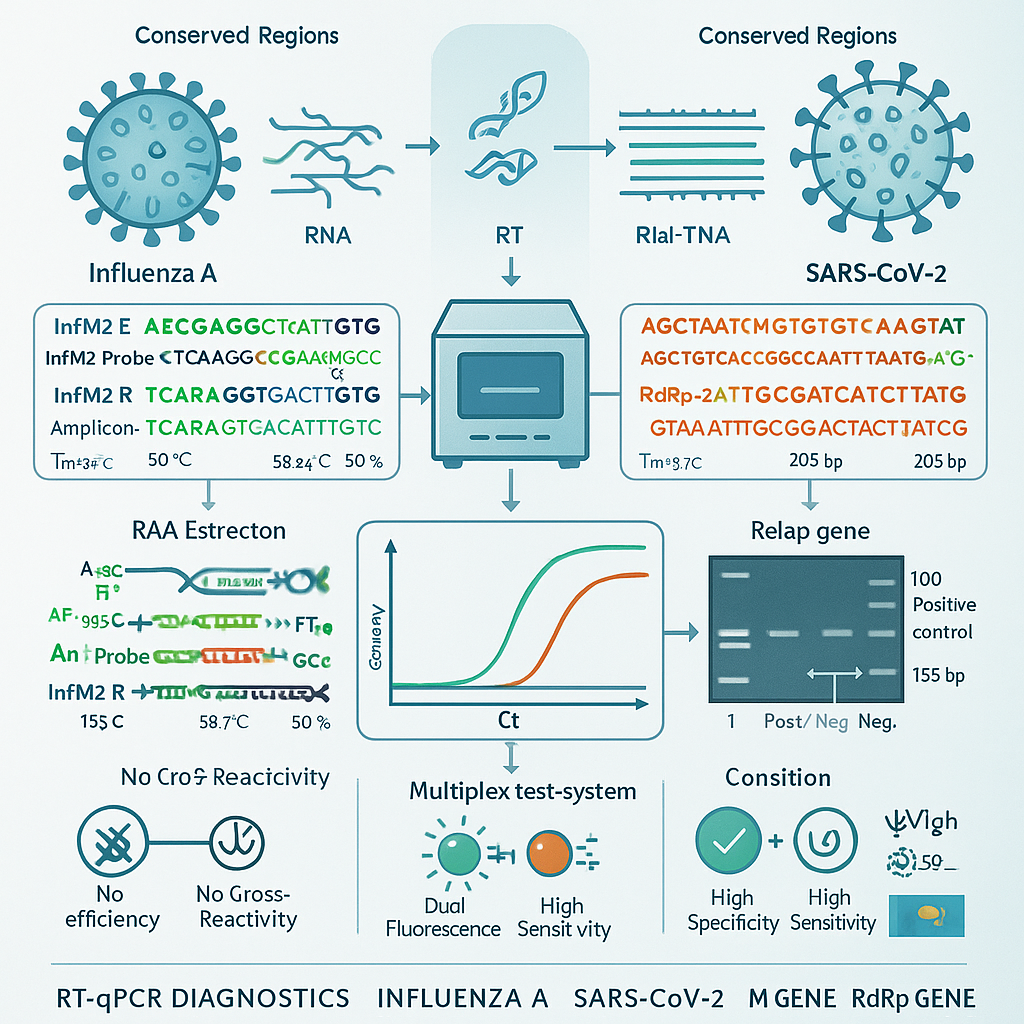
Design of primers and probes for detection of influenza A and SARS-CoV-2 viruses RNA by real-time RT-PCR
DOI:
https://doi.org/10.31489/2026feb1/21-31Keywords:
influenza А, coronavirus, RT-PCR, diagnostics, test-systemAbstract
Cost-effective, accurate, and rapid analysis is essential for testing and diagnosing both common and emerging viruses in clinical virology laboratories. In this study, we designed and selected oligonucleotides for the detection of influenza A virus and SARS-CoV-2 using real-time reverse transcription PCR (RT-qPCR). The development of domestic test systems for diagnosing influenza A virus and SARS-CoV-2 is an urgent task due to the need for early disease detection. The aim of this study is to select primers and probes for the diagnosis of influenza A and SARS-CoV-2 using reverse transcription PCR in real-time (RT-PCR RT). We present the results of designing primers and probes for the identification of influenza A and SARS-CoV-2 RNA. In our studies on the selection of specific primers and probes, the M gene was chosen as a target gene for detecting the influenza A virus, and the RdRp gene for the SARS-CoV-2 virus. A pair of oligonucleotide primers was selected and synthesized for influenza A InfM2 F and InfM2 R, as well as the InfM2 Probe, and for SARS-CoV-2 — RdRp-1 F and RdRp-2 R, the RdRp-2 Probe, which, when performing RT-PCR RT with a working concentration of 20 pmol showed high efficiency in detecting the influenza A virus and SARS-CoV-2. Primers were selected using the Primer Blast and Vector NTI computer programs. The designed primers and probes will be further used to create a domestic multiplex RT-PCR RT test system.
References
1 Припутневич Т. В. Опыт разработки и использования новой тест-системы для скрининга и диагностики инфекций, вызывающих острые респираторные заболевания / Т. В. Припутневич // Эпидемиология и Вакцинопрофилактика. — 2022. — Т. 21. — № 3. — С. 72–79. https://doi.org/10.31631/2073-3046-2022-21-3-72-79 DOI: https://doi.org/10.31631/2073-3046-2022-21-3-72-79
2 Rudan I. Epidemiology and etiology of childhood pneumonia in 2010: estimates of incidence, severe morbidity, mortality, underlying risk factors and causative pathogens for 192 countries / I. Rudan, K. L. O’Brien, H. Nair // J Glob Health. — 2013. — Vol. 3. — No. 1. — P. 10401. DOI: https://doi.org/10.7189/jogh.03.010101
3 Припутневич Т. В. Острые респираторные заболевания и грипп в современном акушерстве: эпидемиологические особенности и проблемы диагностики: обзор литературы / Т. В. Припутневич // Эпидемиология и Вакцинопрофилактика. — 2019. — Т. 18. — № 3. — С. 89–97. https://doi.org/10.31631/2073-3046-2019-18-3-89-97 DOI: https://doi.org/10.31631/2073-3046-2019-18-3-89-97
4 Seto D. S. Acute respiratory infections / D. S. Seto, R. M. Heller // Pediatr Clin North Am. — 1974. — Vol. 21. — No. 3. — P. 683–709. doi: 10.1016/s0031-3955(16)33030-9. PMID: 4369753. DOI: https://doi.org/10.1016/S0031-3955(16)33030-9
5 Орлова Н. В. Острые респираторные заболевания: особенности течения, медикаментозная терапия [Электронный ресурс] / В. Н. Орлова // Медицинский совет. — 2018. — № 15. — Режим доступа: https://cyberleninka.ru/article/n/ostryerespiratornye-zabolevaniya-osobennosti-techeniya-medikamentoznaya-terapiya (Дата обращения: 08.11.2024).
6 Influenza (Seasonal). — 28 February 2025. Site World Health Organization. — Access mode: https://www.who.int/newsroom/fact-sheets/detail/influenza-(seasonal).
7 Ayittey F.K. Economic impacts of Wuhan 2019-nCoV on China and the world / F. K. Ayittey, M. K. Ayittey, N. B. Chiwero, J. S. Kamasah, C. Dzuvor // Journal of medical virology. — 2020. — Vol. 92. — No. 5. — P. 473. https://doi.org/10.1002/jmv.25706 DOI: https://doi.org/10.1002/jmv.25706
8 Википедия: свободная энциклопедия «Пандемия COVID-19» [Электронный ресурс]. — Режим доступа: https://ru.wikipedia.org/wiki/%D0 %9F%D0 %B0 %D0 %BD%D0 %B4 %D0 %B5 %D0 %BC%D0 %B8 %D1 %8F_COVID-19
9 Valencia D. N. Brief review on COVID-19: the 2020 pandemic caused by SARS-CoV-2 / D.N. Valencia // Cureus. — 2020. — Vol. 12. — No. 3. — e7386. doi: 10.7759/cureus.7386. DOI: https://doi.org/10.7759/cureus.7386
10 Ni M. Simultaneous detection and differentiation of SARS-CoV-2, influenza A virus and influenza B virus by one-step quad-ruplex real-time RT-PCR in patients with clinical manifestations / M. Ni, H. Xu, J. Luo, W. Liu, D. Zhou // International Journal of Infectious Diseases. — 2021. — Vol. 103. — P. 517–524. doi: 10.1016/j.ijid.2020.12.027 DOI: https://doi.org/10.1016/j.ijid.2020.12.027
11 Ma L. Development of a Novel Multiplex PCR Method for the Rapid Detection of SARS-CoV-2, Influenza A Virus, and Influenza B Virus / L. Ma, H. Zhu, Y. Jiang, X. Kong, P. Gao, Y. Liu, M. Zhao, G. Deng, Y. Cao // International Journal of Analytical Chemistry. — 2024. — No. 1. — 4950391. doi: 10.1155/2024/4950391 DOI: https://doi.org/10.1155/2024/4950391
12 Bustin S. А. Real-time revers transcription PCR (qRT-PCR) and its potential use in clinical diagnosis / S. A Bustin, R. Mueller // Clinical Science. — 2005. —No. 4. — P. 365–379. DOI: https://doi.org/10.1042/CS20050086
13 COVID-19: Серология, антитела и иммунитет [Электронный ресурс]. — Режим доступа: https://www.who.int/ru/news-room/questions-and-answers/item/coronavirus-disease-covid-19-serology (Дата обращения 08.11.2024)
14 Klumpp-Thomas C. Standardization of ELISA protocols for serosurveys of the SARS-CoV-2 pandemic using clinical and at-home blood sampling / C. Klumpp-Thomas, H. Kalish, M. Drew, S. Hunsberger, K. Snead, M. P. Fay // Nature communications. — 2021. — 12. — P. 1–13. DOI: https://doi.org/10.1038/s41467-020-20383-x
15 Sit T. H. C. et al. Infection of dogs with SARS-CoV-2 / T. H.C. Sit et al. // Nature. — 2020. — Vol. 586. — No. 7831. — P. 776–778. doi: 10.1038/s41586-020-2334-5. DOI: https://doi.org/10.1038/s41586-020-2334-5
16 Gavin P. J. Review of Rapid Diagnostic Tests for Influenza / P. J. Gavin, R. B. Thomson // Clinical and Applied Immunology Reviews. — 2004. — 4(3). — P. 151-172. https://doi.org/10.1016/S1529-1049(03)00064-3. DOI: https://doi.org/10.1016/S1529-1049(03)00064-3
17 Quinlivan M. Comparison of Sensivities of Virus Isolation, Antigen Detection, and Nucleic Acid Amplification for Detection of Equine Influenza Virus / M. Quinlivan, A. Cullinane, M. Nelly // Journal of Clinical Microbiology. — 2004. — Vol. 42. — P. 759–763. doi: 10.1128/JCM.42.2.759-763.2004. DOI: https://doi.org/10.1128/JCM.42.2.759-763.2004
18 Hoffmann E. et al. Universal primer set for the full-length amplification of all influenza A viruses / E. Hoffmann et al. // Archives of Virology. — 2001. — Vol. 146(12). — P. 2275–2289. doi:10.1007/s007050170002 DOI: https://doi.org/10.1007/s007050170002
19 Stone B. et al. Rapid detection and simultaneous subtype differentiation of influenza A viruses by real time PCR / B. Stone et al. // Journal of virological methods. — 2004. — Vol. 117(2). — P. 103–112. doi:10.1016/j.jviromet.2003.12.005 DOI: https://doi.org/10.1016/j.jviromet.2003.12.005
20 Yanagita M. et al. Development of a Real-Time Reverse Transcription PCR Assay System for Detection of Three Subtypes of Influenza A and Influenza B Viruses / M. Yanagita et al. // Rinsho byori. The Japanese Journal of Clinical Pathology. — 2015. — Vol. 63(12). — P. 1365–1370. Japanese. PMID: 27089652.
21 Spackman E. et al. Development of a Real-Time Reverse Transcriptase PCR Assay for Type A Influenza Virus and the Avian H5 and H7 Hemagglutinin Subtypes / E. Spackman et al. // Journal of Clinical Microbiology. — 2002. — Vol. 40(9). — P. 3256-3260. https://doi.org/10.1128/JCM.40.9.3256-3260.2002 DOI: https://doi.org/10.1128/JCM.40.9.3256-3260.2002
22 World Health Organization. CDC protocol of real-time RT-PCR for influenza A(H1N1). — 2009. — [Electronic resource]. — Retrieved from https://www.ecphf.cn/upload/RealtimeRTPCR_SwineH1Assay-2009_20090430.pdf?utm_source
23 Corman V. M. et al. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR / V. M. Corman et al. // Eurosurveillance. — 2020. — Vol. 25(3). — P. 2000045. https://doi.org/10.2807/1560-7917.ES.2020.25.3.2000045 DOI: https://doi.org/10.2807/1560-7917.ES.2020.25.21.2001035
24 Vogels C. B. F. Analytical sensitivity and efficiency comparisons of SARS-CoV-2 qRT-PCR primer-probe sets / C. B. F. Vogels // Nature Microbiology. — 2020. — Vol. 5. — P. 1299–1305. https://doi.org/10.1038/s41564-020-0761-6 DOI: https://doi.org/10.1038/s41564-020-0761-6
25 Dieffenbach C. W. et al. General concepts for PCR primer design / C. W. Dieffenbach et al. // PCR Methods Appl. — 1993. — Vol. 3(3). — P. 30–37. doi: 10.1101/gr.3.3.s30. PMID: 8118394. DOI: https://doi.org/10.1101/gr.3.3.S30